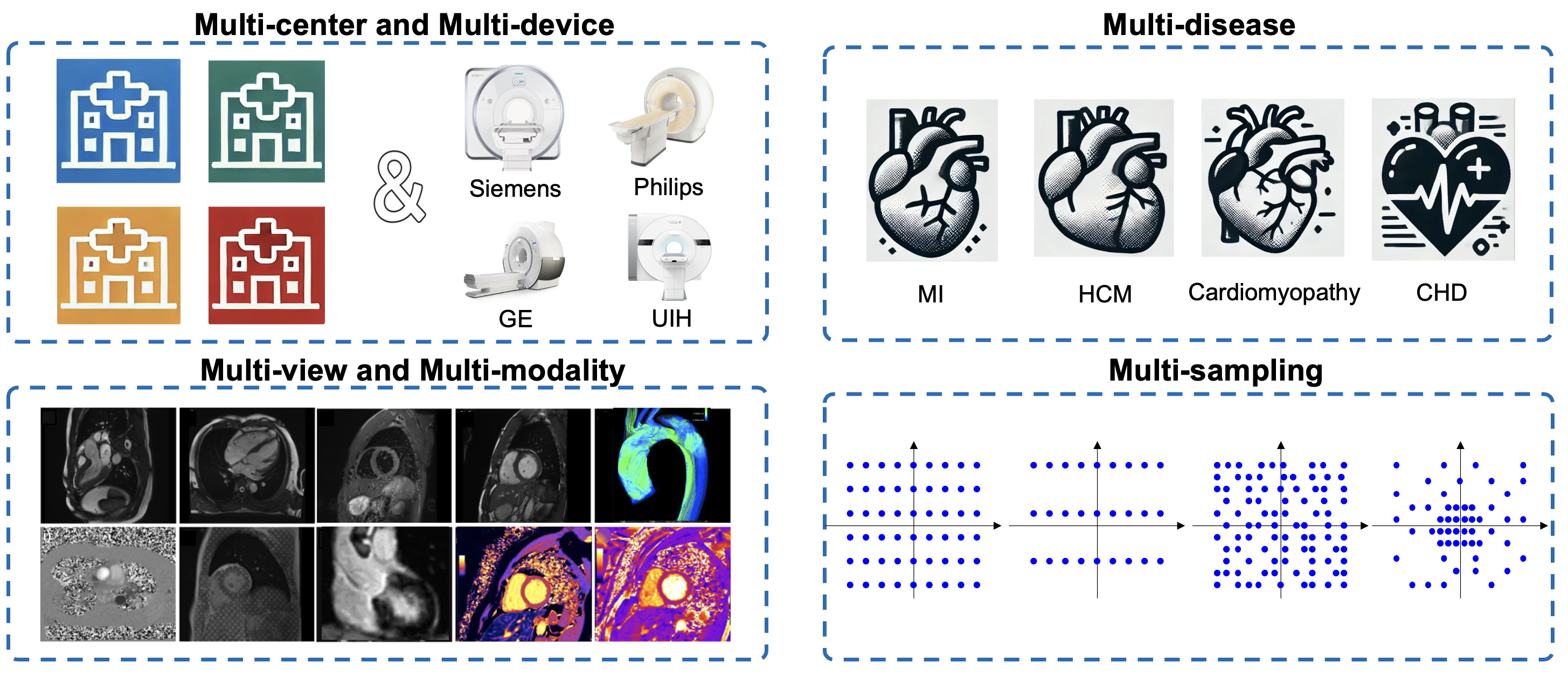
Multi-channel k-space data from
Total of 600 volunteers with multi-parametric CMR imaging. The dataset is seperated into:
The raw k-space data exported from the scanner will be processed and transformed to '.mat' format using the
script provided by each vendor. A readme file will be provided to describe the content and usage of the
data.
Details of raw data description for each modality and view
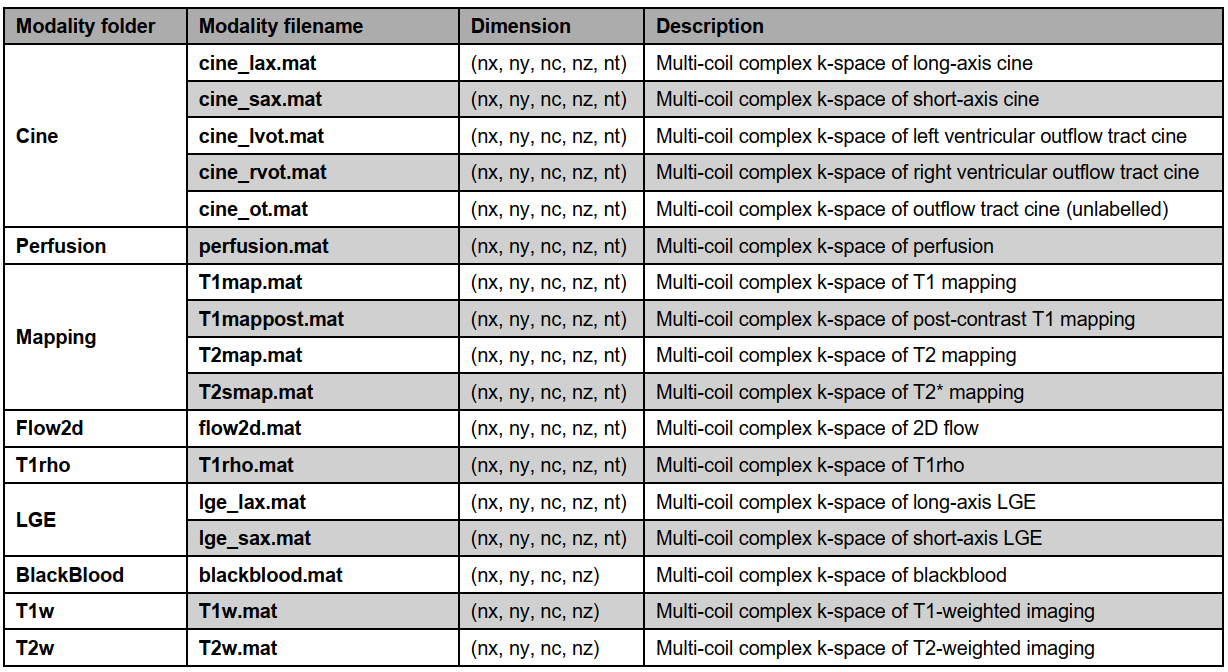
Note: Undersampled and the fully sampled data share the same dimension. nx: matrix size in x-axis (k-space); ny: matrix size in y-axis (k-space); nc: coil array number; nz: slice number (for SAX) or slice group (for LAX, i.e., 3CH, 2CH, and 4CH views); nt: temporal/parametric frame.
Details of different mask types and acceleration factors
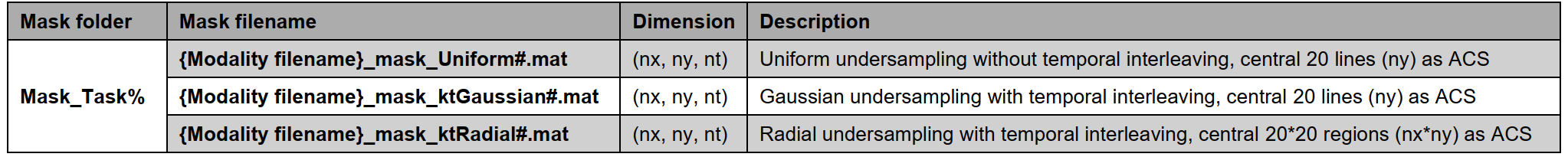
Note: % represents the task number (1 or 2). # represents varied acceleration factors (8x, 16x, 24x), ACS not included for calculations. nx: matrix size in x-axis (k-space); ny: matrix size in y-axis (k-space); nt: temporal/parametric frame. If the data has no temporal/parametric dimension, the corresponding mask dimension becomes (nx, ny).
Taking multi-coil long-axis (LAX) cine in Task
1 as an example, the format of data is as follows:
1) Data in folder 'FullSample': cine_lax.mat
# cine with long-axis view (including 3CH, 2CH, and 4CH views within the nz dimension).
# variable name:
# 'kspace' for fully sampled kspace
# data type: complex kspace data with the dimensions (nx,ny,nc,nz,nt)
-nx: matrix size in x-axis (kspace)
-ny: matrix size in y-axis (kspace)
-nc: coil array number (compressed to 10)
-nz: slice number (for SAX) or slice group (for LAX, i.e., 3CH, 2CH, and 4CH views)
-nt: temporal/parametric frame
2) Data in folder 'UnderSample_Task1':
cine_lax_kus_xxx.mat
# cine with long-axis view (including 3CH, 2CH, and 4CH views within the nz dimension).
# 'xxx' is the corresponding undersampling scenarios. For example, 'ktUniform8' means uniform
undersampling with temporal interleaving at acceleration factor 8x (ACS not included for calculations)
# variable name:
# 'kus' for undersampled kspace
# data type: complex kspace data with the dimensions (nx,ny,nc,nz,nt), the central 20 lines (ny) are
always fully sampled to be used as autocalibration signals (ACS)
-nx: matrix size in x-axis (kspace)
-ny: matrix size in y-axis (kspace)
-nc: coil array number (compressed to 10)
-nz: slice number (for SAX) or slice group (for LAX, i.e., 3CH, 2CH, and 4CH views)
-nt: temporal/parametric frame
3) Data in folder 'Mask_Task1':
cine_lax_mask_xxx.mat
# undersampling mask with long-axis view, the mask is fixed among different nc and nz. But the mask is
interleaved along nt.
# 'xxx' is the corresponding undersampling scenarios. For example, 'ktGaussian8' means Gaussian
undersampling with temporal interleaving at acceleration factor 8x (ACS not included for calculations)
# variable name:
# "mask" for undersampling mask
# data type: 3D binary data with the dimensions (nx,ny,nt), the central 20 lines (ny) or central 20*20
regions (nx*ny) are always fully sampled to be used as autocalibration signals (ACS)
-nx: matrix size in x-axis (kspace)
-ny: matrix size in y-axis (kspace)
-nt: temporal/parametric frame
created with
Website Builder Software .

